Polycistronic Bacterial Plasmid as an Analog of T2A
A polycistronic bacterial plasmid functions as an analog to the T2A system by enabling coexpression of multiple proteins from a single mRNA transcript using multiple ribosome binding sites (RBS) for independent translation initiation. In bacteria, this mirrors the natural operon architecture but contrasts with eukaryotic T2A peptides, which mediate translational skipping within a single open reading frame.
Bacterial Operons: Natural Polycistronic Expression
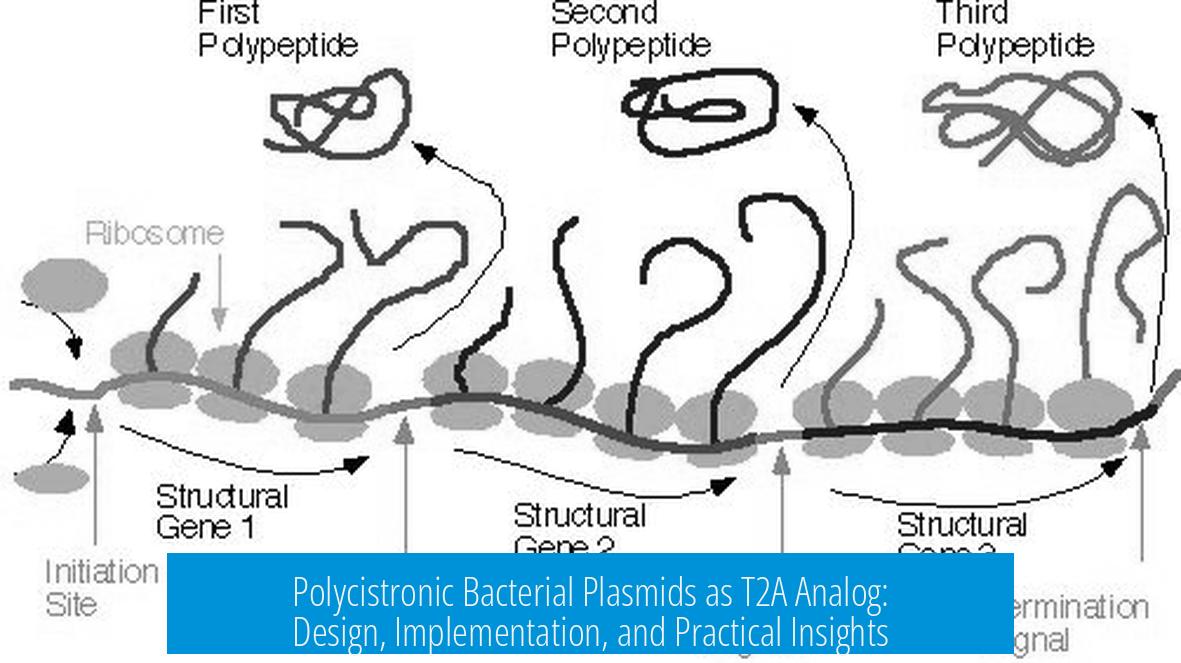
Bacterial genes are often organized into operons. Operons are clusters of related genes controlled by a single promoter, producing one polycistronic mRNA. Each gene in the operon can be translated separately due to multiple ribosomal binding sites. This setup ensures coordinated gene expression and regulation.
Synthetic Polycistronic Plasmids: Design and Implementation
- Promoter and Terminator: Synthetic polycistronic systems employ a strong promoter such as T7 upstream of the gene cluster and a terminator downstream to control transcription.
- Multiple RBS Elements: Each gene is equipped with its own RBS, allowing ribosomes to initiate translation independently. This is crucial for coexpression of separate proteins from the same mRNA.
- Tunable Expression: Synthetic RBS sequences can be designed to have variable strengths. This feature enables precise tuning of protein expression levels in multi-gene constructs.
For example, two genes cloned downstream of a T7 promoter with individual RBS sites will produce one polycistronic mRNA. Upon induction, each protein is translated independently according to its RBS strength.
Practical Considerations for Synthetic Polycistronic Vectors
- Vector Copy Number: Plasmid copy number affects final protein yield and must be matched with experimental needs.
- Induction System: T7 promoters require specific host strains (e.g., T7 Express) and induction conditions (e.g., IPTG). Not all proteins fold well under this system.
- Codon Optimization: Genes should be adapted to the host’s codon usage to maximize translation efficiency.
- Selection Markers: Antibiotic resistance genes maintain plasmid stability in host cells.
Alternative Systems: pET Duet Vector
For coexpression without strict single-transcript demands, the pET Duet vector system offers dual-expression cassettes with individual promoters and RBSs. This system supports simultaneous expression and purification of multi-protein complexes effectively. It is a widely adopted tool in molecular biology applications requiring multiple protein production.
Comparison to Eukaryotic T2A System
| Feature | Bacterial Polycistronic Plasmid | Eukaryotic T2A Peptide |
|---|---|---|
| Transcript | Single polycistronic mRNA with multiple RBS | Single mRNA with one ORF containing T2A sequence |
| Translation Initiation | Independent initiation for each gene via RBS | Ribosome skipping causes cleavage at T2A site, single initiation |
| Protein Products | Separate proteins from separate translation start sites | Separate proteins released from single translation event |
Key Takeaways
- Polycistronic bacterial plasmids mimic natural operon organization for coexpression.
- Multiple synthetic RBS allow independent translation initiation for each protein.
- The system contrasts with T2A’s ribosomal skipping in eukaryotes.
- pET Duet vectors provide flexible coexpression alternatives without a single polycistronic mRNA.
- Codon optimization, vector copy number, and induction conditions affect expression outcomes.
What is the difference between a bacterial polycistronic plasmid and the eukaryotic T2A system?
Bacterial polycistronic plasmids use multiple ribosome binding sites for independent translation of each gene from one mRNA. The T2A system links proteins in one open reading frame, causing ribosome skipping for multiple proteins from a single transcript.
How are protein expression levels controlled in synthetic polycistronic bacterial plasmids?
Each gene has its own synthetic ribosome binding site (RBS) designed with different strengths. This allows tuning each protein’s translation efficiency and adjusting their relative expression levels.
Why might the T7 promoter system not be suitable for all synthetic polycistronic plasmids?
The T7 promoter drives strong expression in specific strains, but it may not work well in all bacterial hosts or expression conditions. Factors like vector copy number and induction conditions also affect protein yield.
What advantages does the pET Duet system offer for coexpression experiments?
The pET Duet system allows coexpression of multiple proteins from separate transcripts within one vector. It is widely used for producing multi-protein complexes without requiring a single polycistronic mRNA.



Leave a Comment